PBPK for Drug-
Drug Interactions
Simulations
Drug-drug interactions (DDIs) can occur when two or more drugs are co-administered, and one drug alters the pharmacokinetics or pharmacodynamics of the other. These interactions can reduce therapeutic efficacy, increase the risk of adverse events, or result in unexpected clinical outcomes. As such, understanding and predicting DDIs is a critical component of drug development and patient safety.
DDIs can arise through a variety of mechanisms:
- CYP enzyme-mediated interactions: Inhibition or induction of cytochrome P450 enzymes can significantly impact drug metabolism, altering plasma levels of the affected drug.
- Non-CYP enzyme-mediated interactions: Other enzymes such as UGTs and MAOs can also be involved in DDIs, particularly for drugs not primarily metabolized by CYPs.
- Transporter-mediated interactions: Membrane transporters such as P-gp, BCRP, OATP, OCTs, and MATEs play key roles in drug absorption, distribution, and excretion. Inhibition or induction of these transporters can lead to altered drug exposure.
- Alteration of gene expression: Some drugs can modulate the expression of metabolic enzymes or transporters, impacting the pharmacokinetics of co-administered compounds.
- Changes in protein binding: Competition for plasma protein binding sites can modify the free (active) concentration of drugs, particularly those that are highly bound.
Due to the potential for DDIs to compromise a drug’s benefit-risk balance, it is essential to predict these interactions early and throughout the drug development process. Regulatory agencies expect detailed, mechanism-based assessments, supported by clinical and modeling data, to be included in drug submissions. Leveraging in silico and in vitro tools to predict and understand DDIs allows for informed risk management and optimized therapeutic strategies.
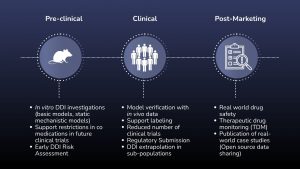
DDIs can be simulated across a range of scenarios, including transporter-mediated interactions, CYP enzyme-mediated mechanisms, drug-gene interactions, non-CYP enzyme pathways, and polymedication cases. These simulations also support regulatory submissions and help assess risks in special populations.

What we can offer
ESQlabs simulates drug-drug interactions and assesses their impact on the pharmacokinetics concomitant agents through PBPK utilizing the OSP suite, PK-Sim and Mobi. Our modeling approaches can quantify the risk of side effects or loss of therapeutic effect due to comedications. The application of PBPK models can be relevant in the early phases of drug development, supporting a rationale characterization of DDI risks as well as used to inform clinical trials, optimize dosing, and help developers and clinicians in their decision-making process to ensure successful drug development and patient safety.
Related Platforms
Large Molecules, Biologics and Novel Modalities
Small Molecules and Chemicals
Related publications and initatives
Meet the Team

Ingrid Michon
Ingrid is a medical biologist with a PhD in biopharmaceutical sciences. After several years in academia, she moved into the pharmaceutical industry, where she built over 12 years of experience in clinical pharmacology, pharmacokinetics and modeling. She also acted as the clinical pharmacology lead for regulatory submissions.
Her passion for PBPK modeling began while working as a clinical pharmacokineticist, assessing DDI liabilities in drug development pipelines. For the past 7 years, she has worked as a PBPK consultant, tackling a wide range of challenges including DDI assessment, first-in-human predictions, pediatric dosing, and regulatory submissions.
Now, at ESQlabs, Ingrid continues to advance 𝗼𝗽𝗲𝗻 𝘀𝗰𝗶𝗲𝗻𝗰𝗲 and help solve complex drug development questions using open-source tools.

Marco Siccardi
Marco is a Clinical Biologist by training with a PhD in molecular pharmacology and PK/PD modelling. He spent over 15 years at the University of Liverpool working on the topic of pharmacogenetics and in developing PBPK approaches for the optimisation of drug delivery, including HIV therapy optimisation.
Marco has most recently been working with CROs in taking this approaches for modelling and simulation approaches and PKTK (Systems Toxicology) models across a number of disease areas.
Marco leads the Systems Toxicology team with the aim to promote collaborative innovation and to develop novel modeling approaches to streamline the toxicological assessment.
- Development of an end-to-end Quantitative Model-Informed Drug Development (MIDD) ECOSYSTEM
- A review of OSP suite PBBM capabilities: looking ahead
- Roadmap for action for advancing aggregate exposure to chemicals in the EU
- Application of High-Throughput PBPK Modeling to Develop an IVIVE Approach for Oral Permeability
- Enhancing PB(P)K Models for the Female Reproductive Tract: A Framework for Local and Systemic Drug Kinetics
- Advancing Maternal-Fetal and Lactation PBK Models for Cross-Species Risk Assessment in Toxicology
- High-Throughput PBPK Framework in R using Open Systems Pharmacology Software for Anti-Tuberculosis Drug Development